
The detection of low VAF variants in cfDNA samples is particularly challenging because high complexity sequencing libraries must be generated using a limited amount of input DNA. The amount of cfDNA ranges from 1 to 100 ng/mL in the plasma, and the allele frequency of tumor DNA in cfDNA is very low, often lower than 1%. 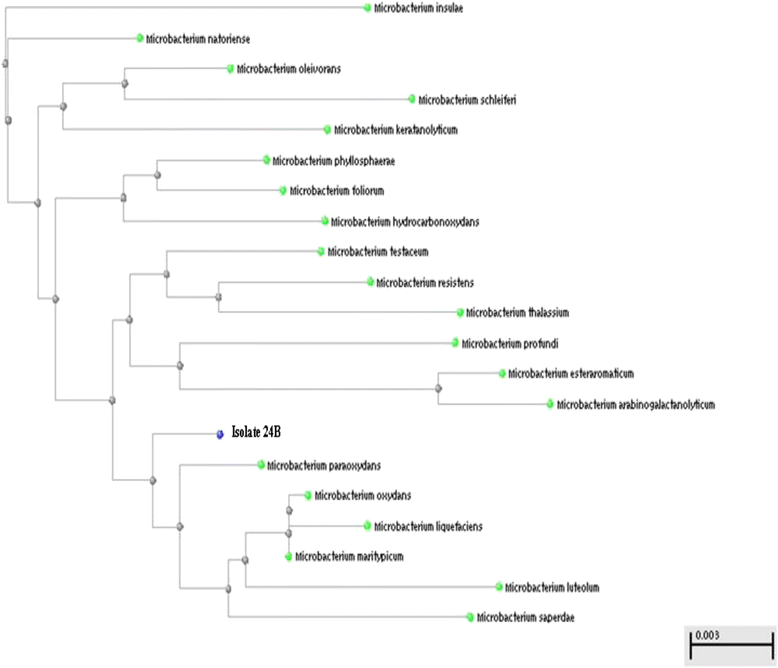
The technical challenge becomes even greater when cancer somatic mutations are interrogated from liquid biopsies such as plasma cell-free DNA (cfDNA) samples. Clinical tissue specimens available for genetic profiling are often minute, which regularly consist of several sections of formalin-fixed paraffin embedded (FFPE) tissues or needle biopsy samples. While the detection of low variant allele fractions (VAFs) requires the sequencing of a sufficient number of molecules, low quantity and quality of DNA extracted from clinical tissue samples often pose obstacles. 
Īs we recently reported, significant proportions of clinically actionable variants have allele fractions as low as less than 5%, often because of low tumor purity, heterogeneity, and secondary tumor driver mutations resulting from treatment. While the targeted sequencing method has been successfully employed for clinical genomic profiling, sufficient sequencing coverage was repeatedly suggested as a prerequisite for the successful implementation of the method in clinical cancer genome profiling. The advantages of targeted deep sequencing are particularly obvious in clinical settings where the selection of therapy is the primary reason for genomic profiling and only a small fraction of identified mutations are potentially responsive to a therapy (i.e., actionable mutations). Whereas whole genome sequencing (WGS) or whole exome sequencing (WES) provides additional information on genomic variants across broad regions of the human genome, targeted sequencing offers distinct advantages over these methods by reducing costs and simplifying data management/analysis. Furthermore, cataloging the most frequently mutated cancer genes across various cancer types has made targeted resequencing an attractive option to cost-effectively analyze genetic alterations in tumor specimens. Because the kits displayed significant variability in different quality metrics, our study offers a practical guideline for researchers to choose appropriate options for PCR-based targeted sequencing and useful benchmark data for evaluating new kits.Ĭancer genome profiling by massively parallel sequencing has rapidly advanced our understanding of the molecular characteristics underlying tumorigenesis. This study is the first systematic evaluation of commercial library construction kits for PCR-based targeted deep sequencing utilizing UMIs. The targeted deep sequencing method based on PCR target enrichment combined with UMI tagging sensitively detected mutations present at a frequency as low as 1% using 6.25 ng of human genomic DNA as the starting material. Based on these comparisons, we selected the Qiagen HASTP for further performance evaluations. We also analyzed the UMIs, including errors, which allowed us to adjust the depth of unique coverage and the length required for sufficient complexity. Regarding the coverage uniformity, the kits from Nugen and NEB performed the best followed by the kits from Qiagen. While the duplicate rates for all kits were dramatically decreased by identifying unique molecules with UMIs, the Qiagen HASTP achieved the highest library complexity based on the depth of unique coverage indicating superb library construction efficiency. We evaluated and compared the performances of the five kits using 50 ng of genomic DNA for the library construction in terms of the library complexity, coverage uniformity, and errors in the UMIs. In this study, we evaluated and compared the performances of five commercial library kits from four vendors: the Archer® Reveal ctDNA™ 28 Kit, NEBNext Direct® Cancer HotSpot Panel, Nugen Ovation® Custom Target Enrichment System, Qiagen Human Comprehensive Cancer Panel(HCCP), and Qiagen Human Actionable Solid Tumor Panel(HASTP). 
Currently, several commercial library construction kits based on PCR enrichment are available for UMIs, but there have been no systematic studies to compare their performances. Recently, this limitation was overcome by assigning a unique molecular identifier(UMI) to each template molecule. Despite the advantages of efficient enrichment, PCR-based methods preclude the identification of PCR duplicates and their subsequent removal. Target enrichment is a critical component of targeted deep next-generation sequencing for the cost-effective and sensitive detection of mutations, which is predominantly performed by either hybrid selection or PCR.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
